API¶
Import vireoSNP as:
import vireoSNP
Read / Load¶
-
vireoSNP.read_cellSNP(dir_name, layers=['AD', 'DP'])[source]¶ Read data from the cellSNP output directory
Parameters: dir_name – directory full path name for cellSNP output Returns: Return type: A disctionary containing AD, DP, cells and variants
-
vireoSNP.read_vartrix(alt_mtx, ref_mtx, cell_file, vcf_file=None)[source]¶ Read data from VarTrix
Parameters: - alt_mtx – sparse matrix file for alternative alleles
- ref_mtx – sparse matrix file for reference alleles
- cell_file – file for cell barcodes, each per line
- vcf_file – the vcf file used for fetch variants in VarTrix
Returns: Return type: A disctionary containing AD, DP, cells and optionally variants
VCF processing¶
Load VCF to matrices¶
-
vireoSNP.vcf.load_VCF(vcf_file, biallelic_only=False, load_sample=True, sparse=True, format_list=None)¶ Initially designed to load VCF from cellSNP output, requiring
- all variants have the same format list;
- a line starting with “#CHROM”, with sample ids.
If these two requirements are satisfied, this function also supports general VCF files, e.g., genotype for multiple samples.
Note, it may take a large memory, please filter the VCF with bcftools first.
Examples
- Load VCF file, e.g., from cellsnp-lite output:
>>> import vireoSNP >>> import numpy as np >>> vcf_dat = vireoSNP.vcf.load_VCF("cellSNP.cells.vcf.gz", sparse=False, >>> biallelic_only=False, format_list=['GT', 'AD', 'DP', 'ALL']) >>> var_ids = np.array(vcf_dat['variants']) >>> samples = np.array(vcf_dat['samples']) >>> GT_mat = np.array(vcf_dat['GenoINFO']['GT']) >>> AD_mat = np.array(vcf_dat['GenoINFO']['AD']).astype(float) >>> DP_mat = np.array(vcf_dat['GenoINFO']['DP']).astype(float) >>> ALL_bases_mat = np.array(vcf_dat['GenoINFO']['ALL'])
Parse genotype probablity to tenseor¶
-
vireoSNP.vcf.parse_donor_GPb(GT_dat, tag='GT', min_prob=0.0)¶ Parse the donor genotype probability tag: GT, GP, or PL
Examples
>>> GProb_tensor = vireoSNP.vcf.parse_donor_GPb(vcf_dat['GenoINFO']['GT'], 'GT')
-
vireoSNP.vcf.match_VCF_samples(VCF_file1, VCF_file2, GT_tag1, GT_tag2)¶ Match donors in two VCF files. Please subset the VCF with bcftools first, as it is more computationally efficient:
bcftools view large_file.vcf.gz -R small_file.vcf.gz -Oz -o sub.vcf.gz
Parameters: - VCF_file1 (str) – the full path of first VCF file, in plain text or gzip / bgzip
- VCF_file2 (str) – the full path of second VCF file, in plain text or gzip / bgzip
- GT_tag1 (str) – the tag for extracting the genotype probability in VCF1: GT, GP, PL
- GT_tag2 (str) – the tag for extracting the genotype probability in VCF2: GT, GP, PL
Plotting¶
Heatmap plot¶
-
vireoSNP.plot.heat_matrix(X, yticks=None, xticks=None, rotation=45, cmap='BuGn', alpha=0.6, display_value=True, row_sort=False, aspect='auto', interpolation='none', **kwargs)[source]¶ Plot heatmap of distance matrix
Parameters: - X (numpy.array or matrix) – The matrix to plot in heatmap
- yticks (list) – The ticks ids for y axis
- xticks (list) – The ticks ids for x axis
- ratation (scalar) – The ratation angel for xticks
- cmap (str) – The colormap for the heatmap, more options: https://matplotlib.org/stable/tutorials/colors/colormaps.html
- alpha (scalar) – The transparency, value between 0 and 1
- display_value (bool) – If True, dispaly the values in the heatmap
- raw_sort (bool) – If True, sort the rows with row index as row_idx = np.argsort(np.dot(X, 2**np.arange(X.shape[1])))
- aspect (str) – aspect in plt.imshow
- interpolation (str) – interpolation in plt.imshow
- **kwargs (keywords & values) – **kwargs for plt.imshow
Returns: Return type: The return from plt.imshow
Examples
>>> from vireoSNP.plot import heat_matrix >>> import numpy as np >>> np.random.seed(1) >>> X = np.random.rand(5, 7) >>> heat_matrix(X)
(Source code, png)
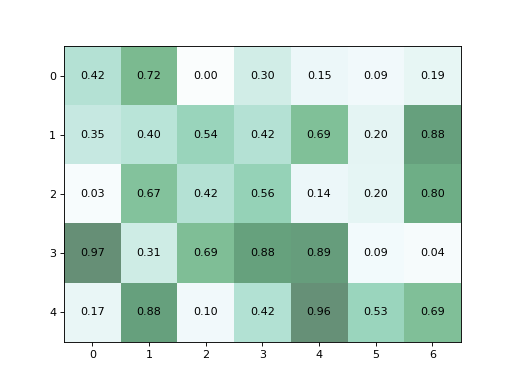
Annotated heatmap plot¶
-
vireoSNP.plot.anno_heat(X, row_anno=None, col_anno=None, row_order_ids=None, col_order_ids=None, xticklabels=False, yticklabels=False, row_cluster=False, col_cluster=False, **kwargs)[source]¶ Heatmap with column or row annotations. Based on seaborn.clustermap() Row or column will be ordered by the annotation group.
Note, haven’t tested if input both row_anno and col_anno.
Vireo Object¶
Objects of type Vireo allow clustering cells by allelic ratio
-
class
vireoSNP.Vireo(n_cell, n_var, n_donor, n_GT=3, learn_GT=True, learn_theta=True, ASE_mode=False, fix_beta_sum=False, beta_mu_init=None, beta_sum_init=None, ID_prob_init=None, GT_prob_init=None)[source]¶ Viroe model: Variational Inference for reconstruction of ensemble origin
The prior can be set via set_prior() before fitting the model.
- beta_mu: numpy array (1, n_GT) or (n_var, n_GT)
- Beta mean parameter of theta’s posterior
- beta_sum: numpy array (1, n_GT) or (n_var, n_GT), same as beta_mu
- Beta concetration parameter of theta’s posterior
- ID_prob: numpy array (n_cell, n_donor)
- Posterior cell assignment probability to each donor
- GT_prob: numpy array (n_var, n_donor, n_GT)
- Posterior genotype probability per variant per donor
-
__init__(n_cell, n_var, n_donor, n_GT=3, learn_GT=True, learn_theta=True, ASE_mode=False, fix_beta_sum=False, beta_mu_init=None, beta_sum_init=None, ID_prob_init=None, GT_prob_init=None)[source]¶ Initialise Vireo model
Note, multiple initializations are highly recomended to avoid local optima.
Parameters: - n_cell (int.) – Number of cells
- n_var (int.) – Number of variants
- n_donor (int.) – Number of donors
- n_GT (int.) – Number of genotype categories
- learn_GT (bool.) – Whether updating GT_prob; otherwise using the initial
- ASE_mode (bool.) – Whether setting allelic ratio theta to be variant specific
- fix_beta_sum (bool.) – Whether fixing the concetration parameter of theta’s posterior
- beta_mu_init (numpy array (1, n_GT) or (n_var, n_GT)) – Initial value of beta_mu, the mean parameter of theta
- beta_sum_init (numpy array (1, n_GT) or (n_var, n_GT), same as beta_mu) – Initial value of beta_sum, the concetration parameter of theta
- ID_prob_init (numpy array (n_cell, n_donor)) – Initial value of ID_prob, cell assignment probability to each donor
- GT_prob_init (numpy array (n_var, n_donor, n_GT)) – Initial value of GT_prob, genotype probability per variant and donor
-
fit(AD, DP, max_iter=200, min_iter=5, epsilon_conv=0.01, delay_fit_theta=0, verbose=True, n_inits=50, nproc=1)[source]¶ Fit Vireo model with coordinate ascent
Parameters: - AD (scipy.sparse.csc_matrix (n_var, n_cell)) – Sparse count matrix for alternative allele
- DP (scipy.sparse.csc_matrix (n_var, n_cell)) – Sparse count matrix for depths, alternative + refeerence alleles
- max_iter (int) – Maximum number of iterations
- min_iter – Minimum number of iterations
- epsilon_conv (float) – Threshold for detecting convergence
- delay_fit_theta (int) – Number of steps to delay updating theta. This can be very useful for common genetics when there is good prior on allelic ratio.
- verbose (bool) – Whether print out log info
BinomMixtureVB Object¶
Objects of type BinomMixtureVB for clustering with binomial
mixture model
-
class
vireoSNP.BinomMixtureVB(n_cell, n_var, n_donor, fix_beta_sum=False, beta_mu_init=None, beta_sum_init=None, ID_prob_init=None)[source]¶ Binomial mixture model with variational inference
The prior can be set via set_prior() before fitting the model.
- beta_mu: numpy array (n_var, n_donor)
- Beta mean parameter of theta’s posterior
- beta_sum: numpy array (n_var, n_donor)
- Beta concetration parameter of theta’s posterior
- ID_prob: numpy array (n_cell, n_donor)
- Posterior cell assignment probability to each donor
-
__init__(n_cell, n_var, n_donor, fix_beta_sum=False, beta_mu_init=None, beta_sum_init=None, ID_prob_init=None)[source]¶ Initialise Vireo model
Note, multiple initializations are highly recomended to avoid local optima.
Parameters: - n_cell (int.) – Number of cells
- n_var (int.) – Number of variants
- n_donor (int.) – Number of donors
- fix_beta_sum (bool.) – Whether fixing the concetration parameter of theta’s posterior
- beta_mu_init (numpy array (n_var, n_donor)) – Initial value of beta_mu, the mean parameter of theta
- beta_sum_init (numpy array (n_var, n_donor)) – Initial value of beta_sum, the concetration parameter of theta
- ID_prob_init (numpy array (n_cell, n_donor)) – Initial value of ID_prob, cell assignment probability to each donor
-
fit(AD, DP, n_init=10, max_iter=200, max_iter_pre=100, random_seed=None, **kwargs)[source]¶ Fit VB with multiple initializations
Parameters: - AD (scipy.sparse.csc_matrix (n_var, n_cell)) – Sparse count matrix for alternative allele
- DP (scipy.sparse.csc_matrix (n_var, n_cell)) – Sparse count matrix for depths, alternative + refeerence alleles
- n_inits (int) – Number of random initialisations to use
- max_iter (int) – Maximum number of iterations for _fit_BV() in best initial
- max_iter_pre (int) – Maximum number of iterations for _fit_BV() in multiple initials
- min_iter – Minimum number of iterations for _fit_BV()
- epsilon_conv (float) – Threshold for detecting convergence for _fit_BV()
- verbose (bool) – Whether print out log info for _fit_BV()
- random_seed (None or int) – Random seed in numpy.random for multiple initializations